Drift Diffusion Model
Random Walk to Full Model
2025
STEP 1: Pure Random Walk - The Foundation
Mathematical Formulation
Discrete Time Random Walk: \[X_n = X_{n-1} + \epsilon_n\]
Where:
- \(X_n\) = position at step \(n\)
- \(X_0 = 0\) (starting position)
- \(\epsilon_n \sim \mathcal{N}(0, \sigma^2)\) (random increment)
Direct Simulation Code
# ==============================================================================
# STEP 1: PURE RANDOM WALK - NO DRIFT, NO BOUNDARIES
# ==============================================================================
# PARAMETERS
n_steps <- 500 # Number of steps to simulate
sigma <- 3 # Standard deviation of noise
# SIMULATION
# set.seed(123)
X <- numeric(n_steps + 1) # Pre-allocate vector
X[1] <- 0 # Starting position
# Generate the random walk step by step
for (n in 2:(n_steps + 1)) {
epsilon <- rnorm(1, mean = 0, sd = sigma) # Random increment
X[n] <- X[n-1] + epsilon # Update position
}
# VISUALIZATION
steps <- 0:n_steps
plot(steps, X, type = "l", col = "steelblue", lwd = 2,
main = "Step 1: Pure Random Walk",
xlab = "Step (n)", ylab = "Position X(n)")
abline(h = 0, lty = 2, col = "gray")
grid()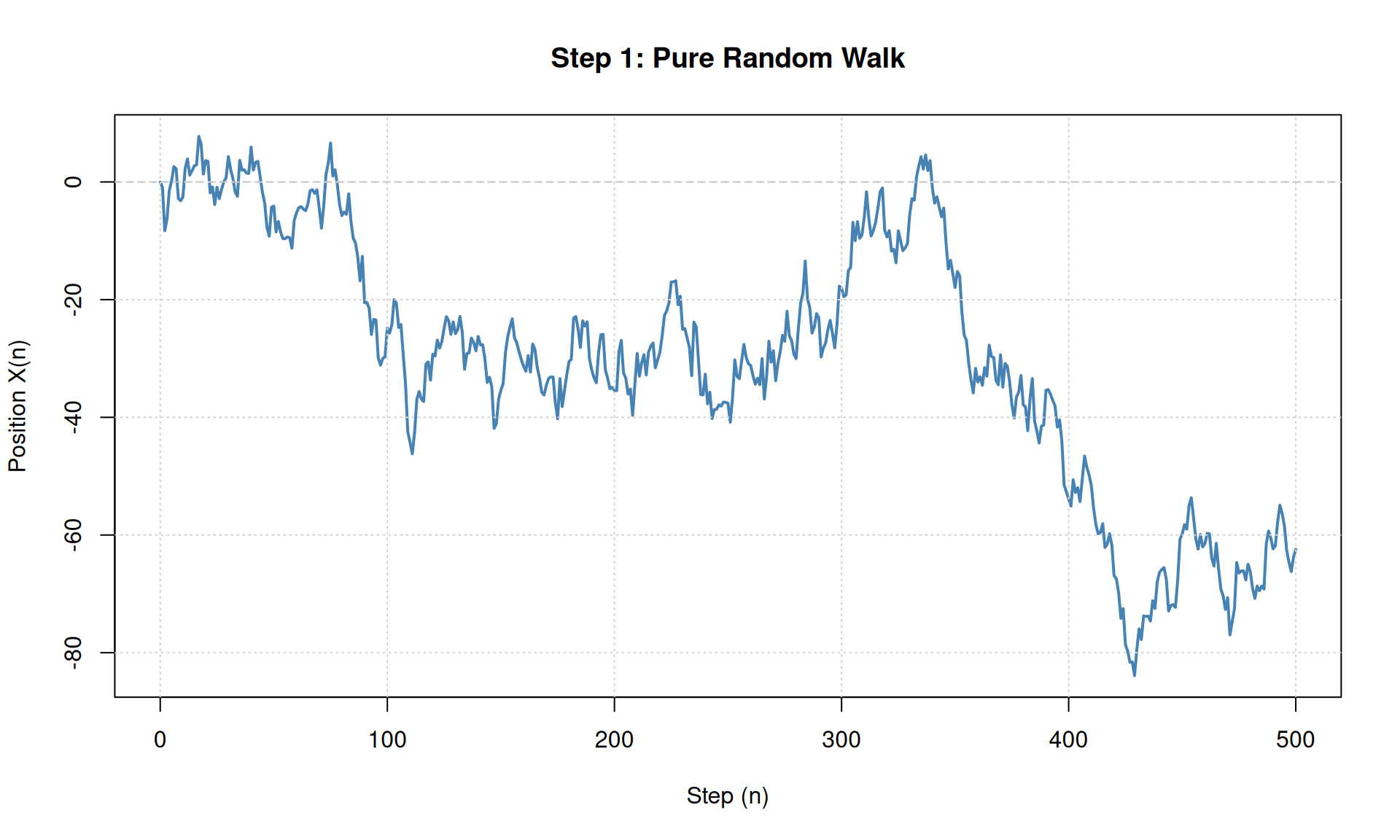
# KEY OBSERVATION: The walk has no direction - just random noise
cat("Final position:", X[n_steps + 1], "\n")## Final position: -62.44304Multiple Paths to See Variability
# Simulate multiple random walks to see the pattern
n_paths <- 10
n_steps <- 500
# Store all paths in a matrix
all_paths <- matrix(0, nrow = n_steps + 1, ncol = n_paths)
set.seed(456)
for (path_id in 1:n_paths) {
X <- numeric(n_steps + 1)
X[1] <- 0
for (n in 2:(n_steps + 1)) {
epsilon <- rnorm(1, mean = 0, sd = 1)
X[n] <- X[n-1] + epsilon
}
all_paths[, path_id] <- X
}
# Plot all paths
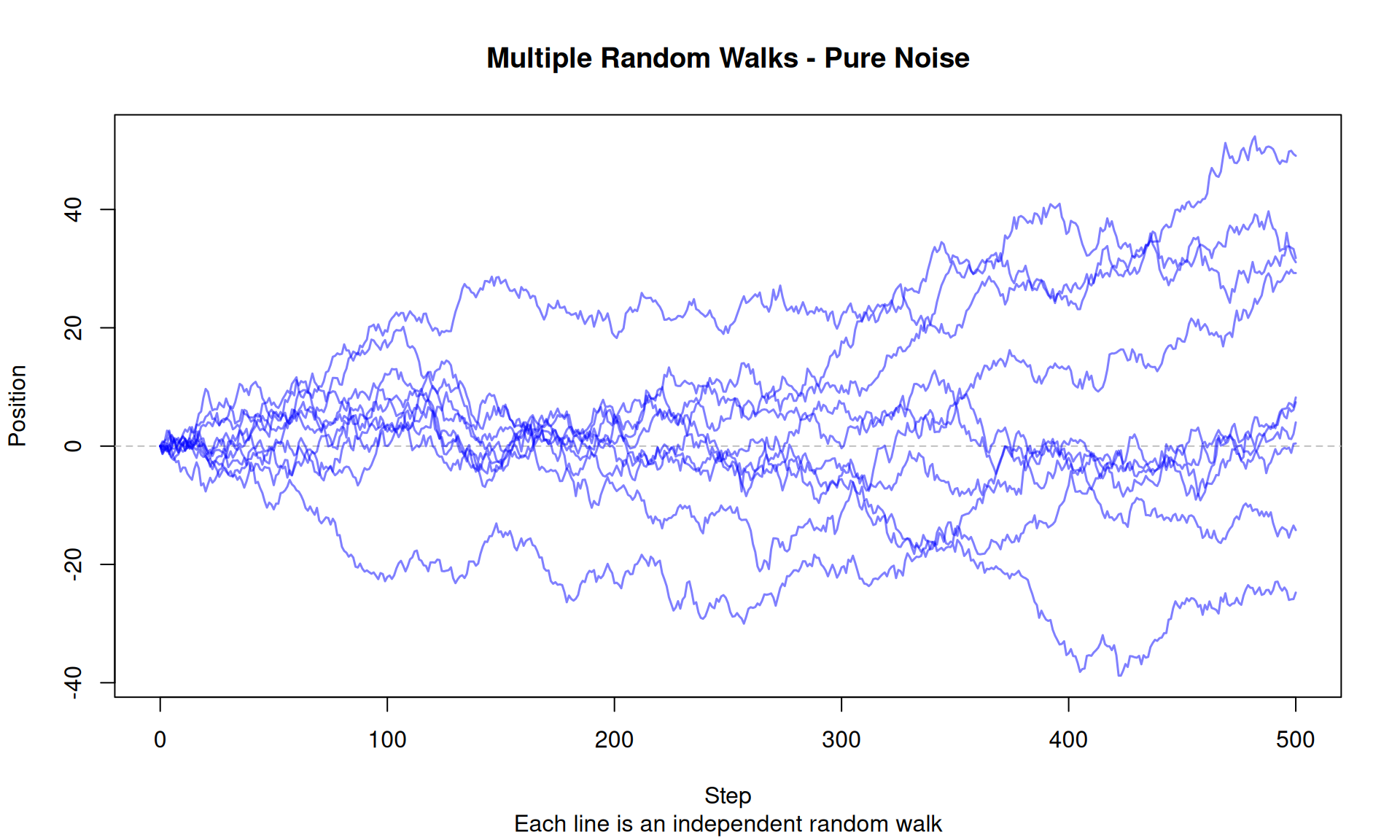
plot(0:n_steps, all_paths[, 1], type = "n",
ylim = range(all_paths),
main = "Multiple Random Walks - Pure Noise",
xlab = "Step", ylab = "Position",
sub = "Each line is an independent random walk")
abline(h = 0, lty = 2, col = "gray")
for (i in 1:n_paths) {
lines(0:n_steps, all_paths[, i], col = rgb(0, 0, 1, 0.5), lwd = 1.5)
}
STEP 2: Adding Drift - Systematic Tendency
Mathematical Formulation
Random Walk with Drift: \[X_n = X_{n-1} + \mu + \epsilon_n\]
Where:
- \(\mu\) = drift parameter (systematic tendency per step)
- Positive \(\mu\) → upward tendency
- Negative \(\mu\) → downward tendency
Direct Simulation with Drift
# ==============================================================================
# STEP 2: RANDOM WALK WITH DRIFT
# ==============================================================================
# PARAMETERS
n_steps <- 500
mu <- -0.1 # Drift parameter
sigma <- 1 # Noise standard deviation
# SIMULATION
# set.seed(789)
X <- numeric(n_steps + 1)
X[1] <- 0
for (n in 2:(n_steps + 1)) {
epsilon <- rnorm(1, mean = 0, sd = sigma)
X[n] <- X[n-1] + mu + epsilon # Now includes drift
}
# VISUALIZATION
plot(0:n_steps, X, type = "l", col = "darkgreen", lwd = 2,
main = paste("Step 2: Random Walk with Drift (μ =", mu, ")"),
xlab = "Step", ylab = "Position")
abline(h = 0, lty = 2, col = "gray")
abline(0, mu, col = "red", lwd = 2, lty = 2) # Expected trajectory
legend("topleft",
legend = c("Actual path", "Expected path (μ × n)"),
col = c("darkgreen", "red"), lwd = 2, lty = c(1, 2))
grid()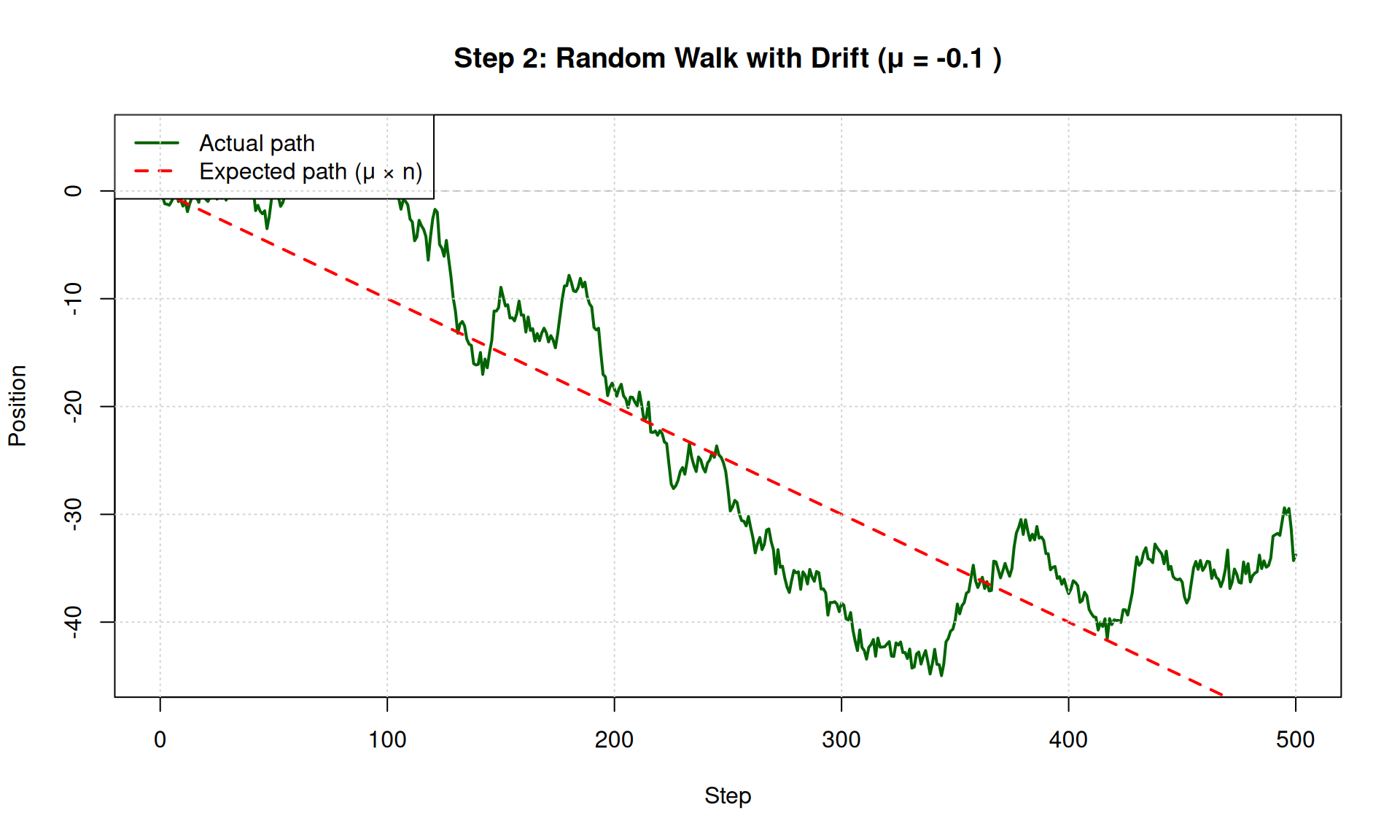
## Starting position: 0## Final position: -33.7729## Expected final position (μ × n): -50Compare Different Drift Values
# Compare mu = -0.2, 0, +0.2
drift_values <- c(-0.2, 0, 0.2)
n_steps <- 500
colors <- c("red", "gray", "blue")
# Setup plot
plot(0:n_steps, 0:n_steps, type = "n",
ylim = c(-150, 150),
main = "Effect of Drift Parameter",
xlab = "Step", ylab = "Position")
abline(h = 0, lty = 2)
grid()
set.seed(100)
for (i in 1:3) {
mu <- drift_values[i]
X <- numeric(n_steps + 1)
X[1] <- 0
for (n in 2:(n_steps + 1)) {
epsilon <- rnorm(1, mean = 0, sd = 1)
X[n] <- X[n-1] + mu + epsilon
}
lines(0:n_steps, X, col = colors[i], lwd = 2)
}
legend("topleft",
legend = paste("μ =", drift_values),
col = colors, lwd = 2)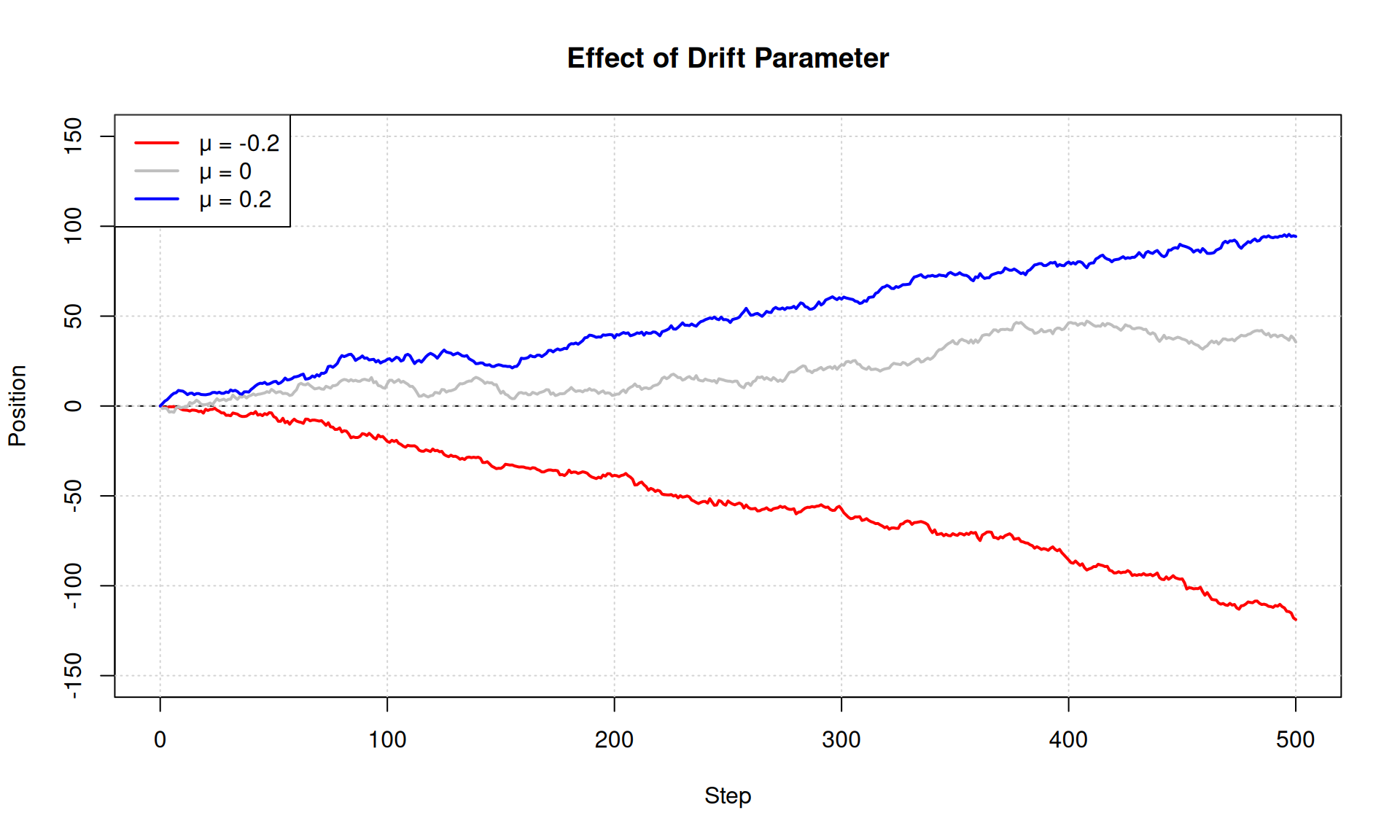
STEP 3: From Discrete to Continuous Time
Mathematical Formulation
Continuous Time Approximation: \[X(t + \Delta t) = X(t) + v \cdot \Delta t + s \cdot \sqrt{\Delta t} \cdot Z\]
Where:
- \(t\) = continuous time (in seconds)
- \(\Delta t\) = time step size (e.g., 0.001 seconds)
- \(v\) = drift rate (evidence per second)
- \(s\) = diffusion coefficient (noise intensity) - \(Z \sim \mathcal{N}(0, 1)\)
KEY: Why \(\sqrt{\Delta t}\)?
- Preserves correct variance scaling in continuous limit
- Ensures Var[noise] = \(s^2 \cdot \Delta t\) per time step
Direct Simulation in Continuous Time
# ==============================================================================
# STEP 3: CONTINUOUS TIME APPROXIMATION (EULER-MARUYAMA)
# ==============================================================================
# PARAMETERS
v <- 0.3 # Drift rate (evidence per second)
s <- 1.0 # Diffusion coefficient (noise intensity)
dt <- 0.001 # Time step (1 millisecond)
T_max <- 2.0 # Maximum time (seconds)
# SIMULATION
n_steps <- round(T_max / dt)
time <- seq(0, T_max, by = dt)
X <- numeric(n_steps + 1)
X[1] <- 0
for (i in 2:(n_steps + 1)) {
Z <- rnorm(1, mean = 0, sd = 1)
X[i] <- X[i-1] + v * dt + s * sqrt(dt) * Z
}
# VISUALIZATION
plot(time, X, type = "l", col = "purple", lwd = 2,
main = "Step 3: Continuous Time Random Walk (Wiener Process)",
xlab = "Time (seconds)", ylab = "Evidence X(t)",
sub = paste("v =", v, ", s =", s, ", Δt =", dt))
abline(h = 0, lty = 2, col = "gray")
abline(0, v, col = "red", lwd = 2, lty = 2)
grid()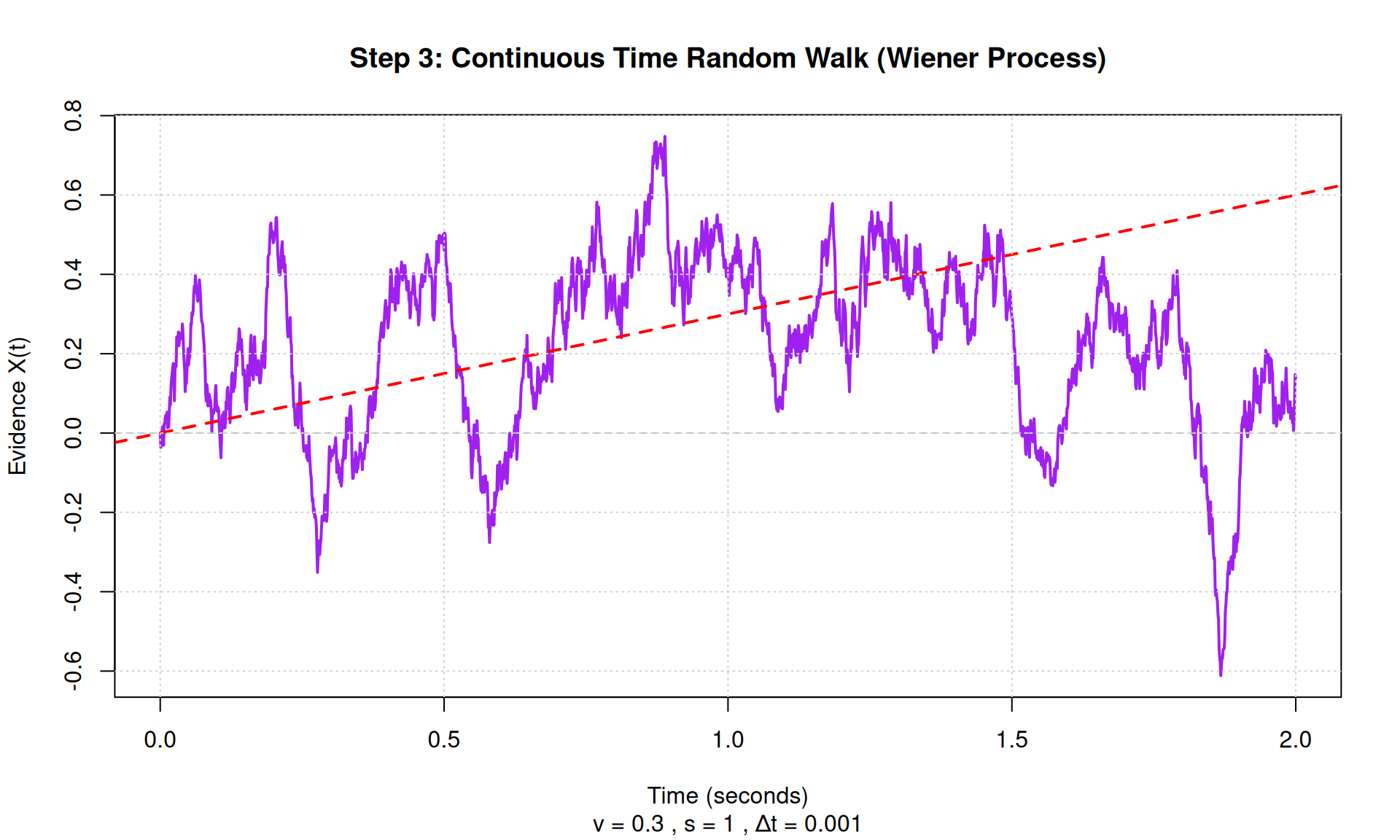
## Total time simulated: 2 seconds## Number of time steps: 2000## Final position: 0.14STEP 4: Adding Decision Boundaries
Mathematical Formulation
Decision Rule:
- Upper boundary at \(X = a\) → Decision “A” (e.g., “yes”, “correct”)
- Lower boundary at \(X = 0\) → Decision “B” (e.g., “no”, “error”)
- Starting point at \(X = z\) (typically \(z = a/2\) for unbiased)
Process:
1. Start at \(X(0) = z\)
2. Accumulate evidence: \(X(t + \Delta t) = X(t) + v \cdot \Delta t + s \cdot \sqrt{\Delta t} \cdot Z\)
3. Stop when \(X(t) \geq a\) OR \(X(t) \leq 0\)
4. Record: choice (which boundary) and RT (time to boundary)
Direct Simulation with Boundaries
# ==============================================================================
# STEP 4A: SEE THE RANDOM WALKS WITH BOUNDARIES
# ==============================================================================
# PARAMETERS
v <- 0.1 # Drift rate (moderate)
a <- 2.0 # Boundary separation
z <- 0.5 * a # Starting point (middle)
s <- 1.0 # Diffusion coefficient
dt <- 0.001 # Time step
max_time <- 2.0 # Maximum time
n_paths <- 15 # Number of paths to show
# Storage for all paths
all_paths <- list()
all_times <- list()
all_choices <- character(n_paths)
all_RTs <- numeric(n_paths)
set.seed(222)
cat("Simulating", n_paths, "visible paths...\n\n")## Simulating 15 visible paths...# Simulate each path
for (path_id in 1:n_paths) {
evidence <- z
time_elapsed <- 0
step_counter <- 0
# Store this path
evidence_path <- evidence
time_path <- 0
# Run until boundary hit
while (evidence > 0 && evidence < a && time_elapsed < max_time) {
Z <- rnorm(1, 0, 1)
evidence <- evidence + v * dt + s * sqrt(dt) * Z
time_elapsed <- time_elapsed + dt
step_counter <- step_counter + 1
# Store every 10th point to keep file size manageable
if (step_counter %% 10 == 0) {
evidence_path <- c(evidence_path, evidence)
time_path <- c(time_path, time_elapsed)
}
}
# Always store the final point where boundary was hit
evidence_path <- c(evidence_path, evidence)
time_path <- c(time_path, time_elapsed)
# Record outcome
if (time_elapsed >= max_time) {
all_choices[path_id] <- "Timeout"
all_RTs[path_id] <- NA
} else {
all_choices[path_id] <- ifelse(evidence >= a, "Upper", "Lower")
all_RTs[path_id] <- time_elapsed
}
# Store path
all_paths[[path_id]] <- evidence_path
all_times[[path_id]] <- time_path
cat("Path", path_id, ":", all_choices[path_id], "boundary, RT =",
round(all_RTs[path_id], 3), "s\n")
}## Path 1 : Timeout boundary, RT = NA s
## Path 2 : Lower boundary, RT = 0.365 s
## Path 3 : Lower boundary, RT = 0.097 s
## Path 4 : Lower boundary, RT = 0.92 s
## Path 5 : Upper boundary, RT = 0.359 s
## Path 6 : Lower boundary, RT = 1.03 s
## Path 7 : Lower boundary, RT = 1.82 s
## Path 8 : Lower boundary, RT = 0.363 s
## Path 9 : Lower boundary, RT = 1.044 s
## Path 10 : Upper boundary, RT = 0.471 s
## Path 11 : Upper boundary, RT = 0.384 s
## Path 12 : Lower boundary, RT = 0.856 s
## Path 13 : Upper boundary, RT = 0.432 s
## Path 14 : Upper boundary, RT = 0.628 s
## Path 15 : Upper boundary, RT = 0.536 s# VISUALIZATION: All paths on one plot
plot(NULL, xlim = c(0, max_time), ylim = c(-0.15, a + 0.15),
main = "Step 4: Evidence Paths with Decision Boundaries",
xlab = "Time (seconds)", ylab = "Evidence",
sub = paste(n_paths, "independent random walks"))
# Draw boundaries FIRST (so paths go on top)
abline(h = a, col = "red", lwd = 3, lty = 1)
abline(h = 0, col = "red", lwd = 3, lty = 1)
abline(h = z, col = "orange", lwd = 2, lty = 2)
# Add labels
text(max_time * 0.98, a + 0.08, "UPPER BOUNDARY", col = "red", adj = 1, font = 2)
text(max_time * 0.98, -0.08, "LOWER BOUNDARY", col = "red", adj = 1, font = 2)
text(0.05, z + 0.08, "START", col = "orange", adj = 0)
# Draw each path
for (i in 1:n_paths) {
# Color based on which boundary was hit
if (all_choices[i] == "Upper") {
path_color <- rgb(0, 0, 0.8, 0.6) # Blue for upper
} else if (all_choices[i] == "Lower") {
path_color <- rgb(0.8, 0, 0, 0.6) # Red for lower
} else {
path_color <- rgb(0.5, 0.5, 0.5, 0.4) # Gray for timeout
}
lines(all_times[[i]], all_paths[[i]], col = path_color, lwd = 1.5)
}
# Add legend
legend("topleft",
legend = c("Upper boundary", "Lower boundary", "Timeout"),
col = c(rgb(0, 0, 0.8, 0.6), rgb(0.8, 0, 0, 0.6), rgb(0.5, 0.5, 0.5, 0.4)),
lwd = 2, bg = "white")
grid()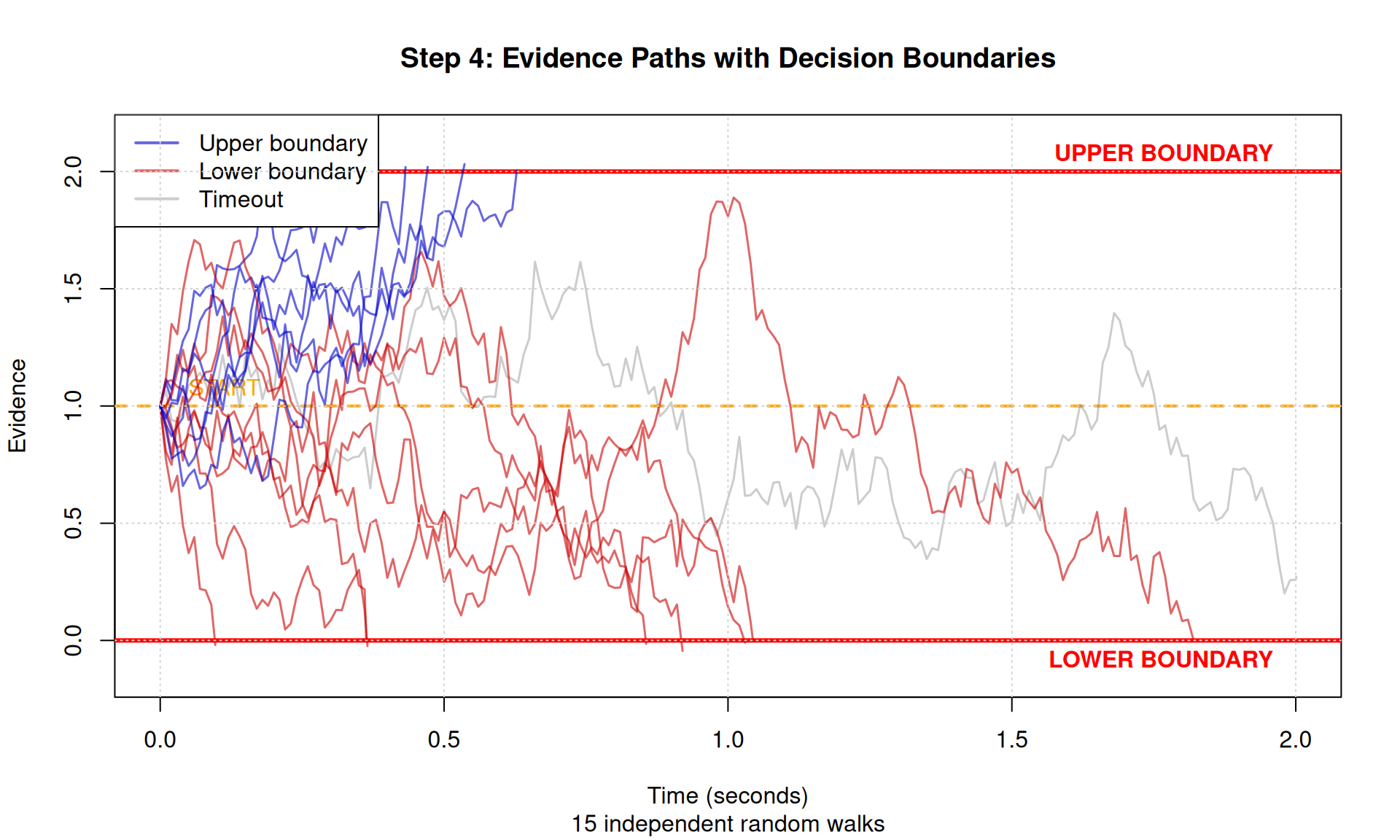
##
## === SUMMARY ===## Upper boundary: 6 / 15## Lower boundary: 8 / 15## Timeouts: 1 / 15Key Observations
Look at the plot above and notice:
- All paths start at the same point (orange line)
- Paths wander randomly but with upward drift (v > 0)
- Blue paths hit upper boundary → these are “correct” responses
- Red paths hit lower boundary → these are “errors”
- Different RTs even for same boundary (randomness!)
- More blue than red because drift favors upper boundary
Simulate Multiple Trials
# Simulate many trials to see distributions
n_trials <- 500
v <- 0.3
a <- 1.0
z <- 0.5 * a
s <- 1.0
dt <- 0.001
max_time <- 5.0
# Store results
choices <- character(n_trials)
RTs <- numeric(n_trials)
set.seed(333)
cat("Simulating", n_trials, "trials...\n")## Simulating 500 trials...for (trial in 1:n_trials) {
evidence <- z
time_elapsed <- 0
while (evidence > 0 && evidence < a && time_elapsed < max_time) {
Z <- rnorm(1, 0, 1)
evidence <- evidence + v * dt + s * sqrt(dt) * Z
time_elapsed <- time_elapsed + dt
}
if (time_elapsed >= max_time) {
choices[trial] <- NA
RTs[trial] <- NA
} else {
choices[trial] <- ifelse(evidence >= a, "Upper", "Lower")
RTs[trial] <- time_elapsed
}
}
# Remove NA trials
valid <- !is.na(choices)
choices <- choices[valid]
RTs <- RTs[valid]
# RESULTS
cat("\nResults from", sum(valid), "valid trials:\n")##
## Results from 500 valid trials:## P(Upper boundary): 0.582## Mean RT: 0.264 seconds## Mean RT (Upper): 0.274 seconds## Mean RT (Lower): 0.251 seconds# VISUALIZATION: RT Distributions
par(mfrow = c(1, 2))
# Histogram by choice
hist(RTs[choices == "Upper"], breaks = 30, col = rgb(0, 0, 1, 0.5),
main = "RT Distributions by Choice",
xlab = "Reaction Time (seconds)", xlim = range(RTs))
hist(RTs[choices == "Lower"], breaks = 30, col = rgb(1, 0, 0, 0.5), add = TRUE)
legend("topright", legend = c("Upper", "Lower"),
fill = c(rgb(0, 0, 1, 0.5), rgb(1, 0, 0, 0.5)))
# Overall RT distribution
hist(RTs, breaks = 30, col = "lightgray",
main = "Overall RT Distribution",
xlab = "Reaction Time (seconds)")
abline(v = mean(RTs), col = "red", lwd = 2)
text(mean(RTs), par("usr")[4] * 0.9,
paste("Mean =", round(mean(RTs), 3)),
col = "red", adj = 0)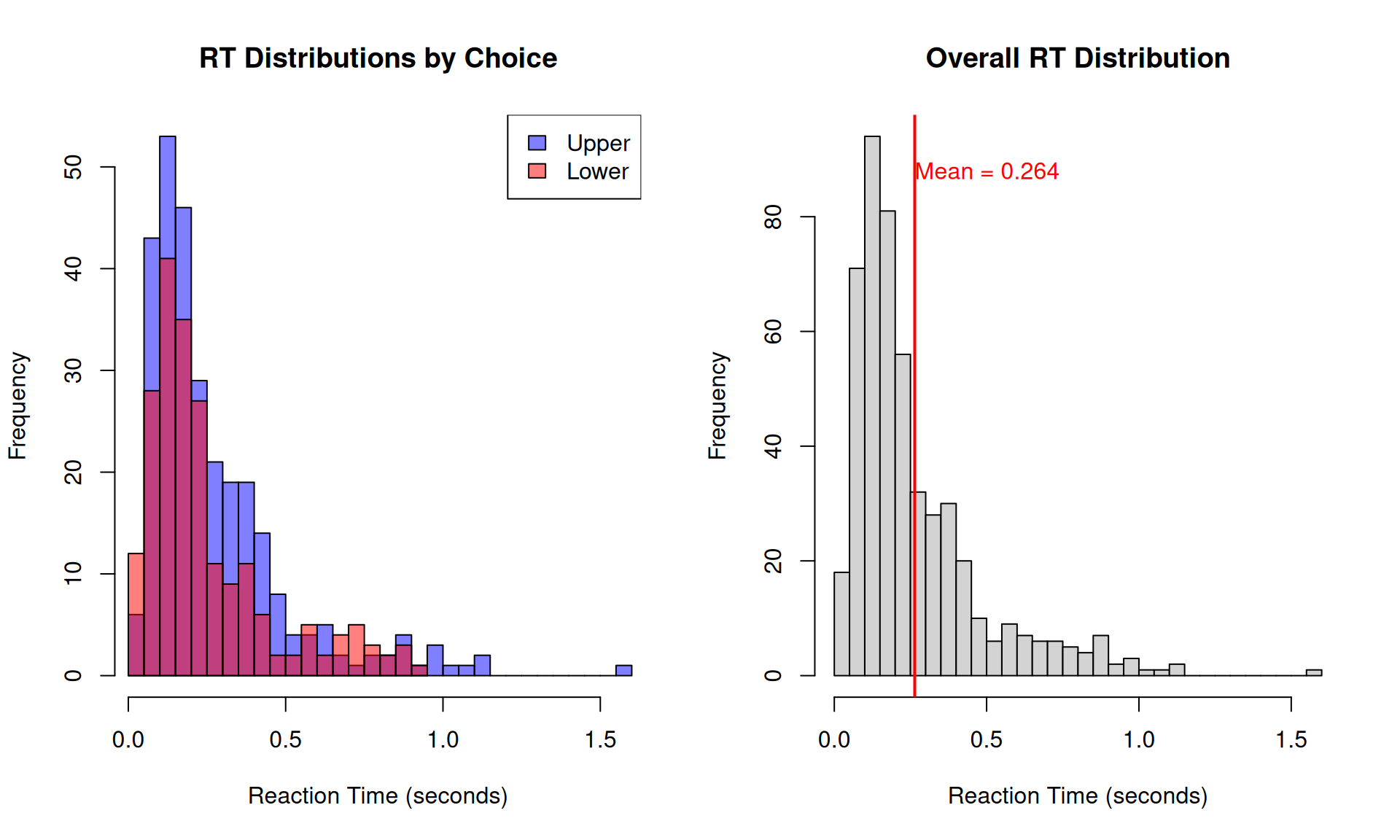
STEP 5: Adding Non-Decision Time
Mathematical Formulation
Total RT = Decision Time + Non-Decision Time
\[RT_{total} = RT_{decision} + T_{er}\]
Where:
- \(RT_{decision}\) = time to reach boundary
- \(T_{er}\) = non-decision time (encoding + motor response)
- Typically \(T_{er}\) = 0.2 - 0.4 seconds
Simulation with Non-Decision Time
# ==============================================================================
# STEP 5: ADDING NON-DECISION TIME
# ==============================================================================
# PARAMETERS
v <- 0.3
a <- 1.0
z <- 0.5 * a
s <- 1.0
Ter <- 0.3 # Non-decision time (NEW!)
dt <- 0.001
max_time <- 5.0
# Simulate one trial
evidence <- z
decision_time <- 0
while (evidence > 0 && evidence < a && decision_time < max_time) {
Z <- rnorm(1, 0, 1)
evidence <- evidence + v * dt + s * sqrt(dt) * Z
decision_time <- decision_time + dt
}
if (decision_time >= max_time) {
choice <- NA
RT_total <- NA
} else {
choice <- ifelse(evidence >= a, "Upper", "Lower")
RT_total <- decision_time + Ter # Add non-decision time!
}
cat("Decision time:", round(decision_time, 3), "seconds\n")## Decision time: 0.232 seconds## Non-decision time: 0.3 seconds## Total RT: 0.532 seconds## Choice: Upper# VISUALIZATION: Components of RT
barplot(c(decision_time, Ter),
names.arg = c("Decision Time", "Non-Decision Time"),
col = c("steelblue", "orange"),
main = "Components of Reaction Time",
ylab = "Time (seconds)",
ylim = c(0, RT_total * 1.2))
text(0.7, decision_time/2, round(decision_time, 3), col = "white")
text(1.9, decision_time + Ter/2, Ter, col = "white")
text(1.3, RT_total + 0.05, paste("Total RT =", round(RT_total, 3)))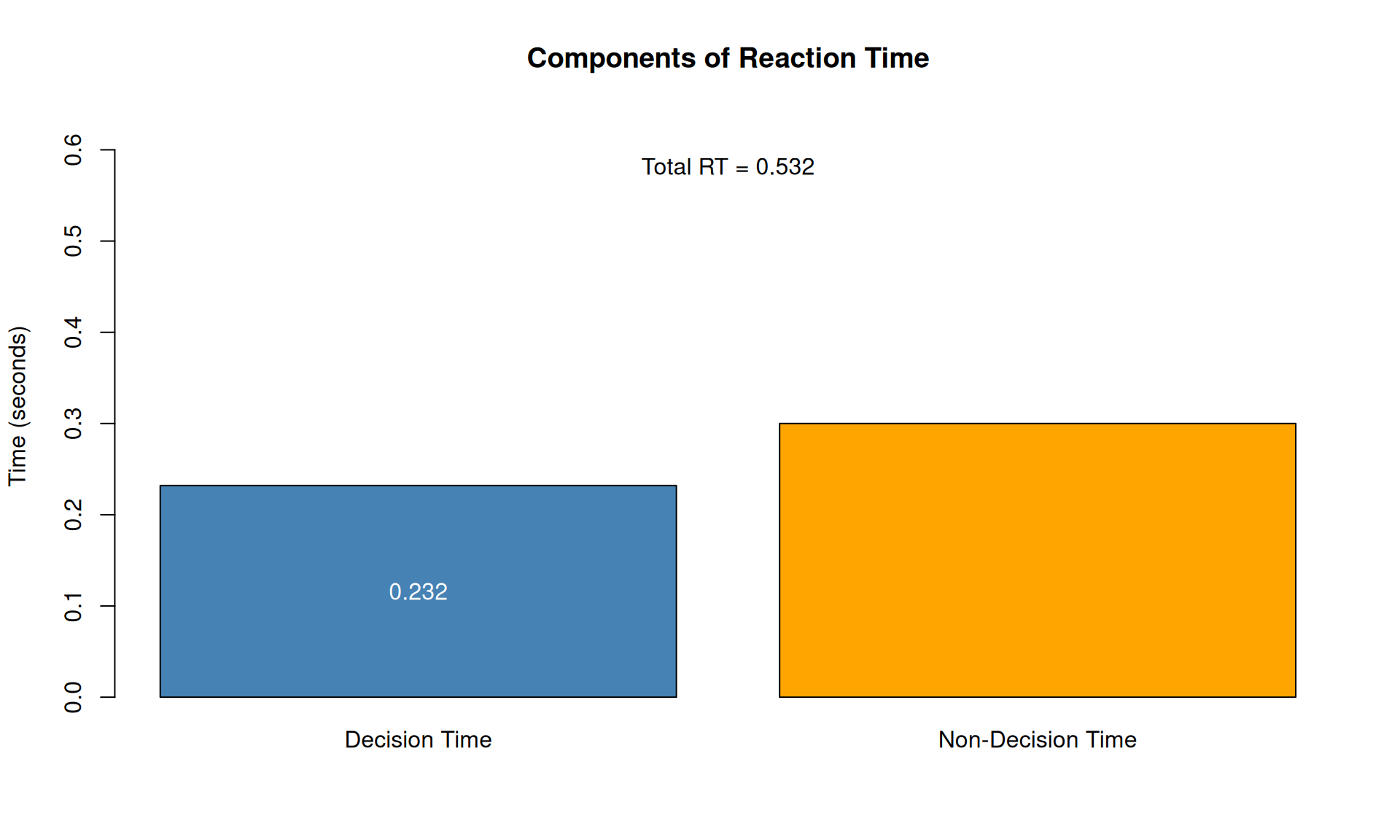
STEP 6: The Complete DDM
Mathematical Formulation
Full DDM Process:
- Initialize: \(X(0) = z\)
- Accumulate: \(X(t + \Delta t) = X(t) + v \cdot \Delta t + s \cdot \sqrt{\Delta t} \cdot Z\)
- Decide: Stop when \(X(t) \geq a\) or \(X(t) \leq 0\)
- Respond: \(RT = t_{decision} + T_{er}\)
Parameters:
- \(v\) = drift rate (quality of evidence)
- \(a\) = boundary separation (speed-accuracy tradeoff)
- \(z\) = starting point (bias)
- \(s\) = diffusion coefficient (noise)
- \(T_{er}\) = non-decision time
Complete DDM Simulation
# ==============================================================================
# STEP 6: COMPLETE DDM - ALL PARAMETERS
# ==============================================================================
# PARAMETERS (All Together!)
v <- 0.25 # Drift rate
a <- 1.2 # Boundary separation
z_prop <- 0.5 # Starting point (as proportion)
z <- z_prop * a # Convert to absolute
s <- 1.0 # Diffusion coefficient
Ter <- 0.25 # Non-decision time
dt <- 0.001 # Time step
max_time <- 10.0 # Max time
# Simulate many trials
n_trials <- 1000
all_choices <- character(n_trials)
all_RTs <- numeric(n_trials)
set.seed(555)
cat("Running complete DDM with", n_trials, "trials...\n\n")## Running complete DDM with 1000 trials...for (trial in 1:n_trials) {
evidence <- z
time <- 0
while (evidence > 0 && evidence < a && time < max_time) {
Z <- rnorm(1, 0, 1)
evidence <- evidence + v * dt + s * sqrt(dt) * Z
time <- time + dt
}
if (time >= max_time) {
all_choices[trial] <- NA
all_RTs[trial] <- NA
} else {
all_choices[trial] <- ifelse(evidence >= a, "Upper", "Lower")
all_RTs[trial] <- time + Ter
}
}
# Clean data
valid <- !is.na(all_choices)
choices <- all_choices[valid]
RTs <- all_RTs[valid]
# SUMMARY STATISTICS
cat("RESULTS:\n")## RESULTS:## Valid trials: 1000 / 1000## Accuracy (P(Upper)): 0.607## Mean RT: 0.631 seconds## Median RT: 0.543 seconds## SD RT: 0.307 seconds## By Choice:cat("Upper - N:", sum(choices == "Upper"),
", Mean RT:", round(mean(RTs[choices == "Upper"]), 3), "\n")## Upper - N: 607 , Mean RT: 0.622cat("Lower - N:", sum(choices == "Lower"),
", Mean RT:", round(mean(RTs[choices == "Lower"]), 3), "\n")## Lower - N: 393 , Mean RT: 0.643# COMPREHENSIVE VISUALIZATION
par(mfrow = c(2, 2))
# 1. RT distributions by choice
hist(RTs[choices == "Upper"], breaks = 40, col = rgb(0, 0, 1, 0.5),
main = "RT Distributions",
xlab = "RT (seconds)", xlim = range(RTs), freq = FALSE)
hist(RTs[choices == "Lower"], breaks = 40, col = rgb(1, 0, 0, 0.5),
add = TRUE, freq = FALSE)
legend("topright", c("Upper", "Lower"),
fill = c(rgb(0, 0, 1, 0.5), rgb(1, 0, 0, 0.5)))
# 2. Choice proportions
choice_counts <- table(choices)
barplot(choice_counts, col = c("salmon", "steelblue"),
main = "Choice Frequencies",
ylab = "Count")
# 3. RT quantiles by choice
quantiles <- c(0.1, 0.3, 0.5, 0.7, 0.9)
q_upper <- quantile(RTs[choices == "Upper"], quantiles)
q_lower <- quantile(RTs[choices == "Lower"], quantiles)
plot(quantiles, q_upper, type = "b", col = "blue", lwd = 2,
ylim = range(c(q_upper, q_lower)),
main = "RT Quantiles by Choice",
xlab = "Quantile", ylab = "RT (seconds)")
lines(quantiles, q_lower, type = "b", col = "red", lwd = 2)
legend("topleft", c("Upper", "Lower"), col = c("blue", "red"), lwd = 2)
# 4. Parameter summary
plot.new()
text(0.5, 0.8, "DDM Parameters", cex = 1.5, font = 2)
text(0.2, 0.6, "Drift rate (v):", adj = 0)
text(0.7, 0.6, v, adj = 0)
text(0.2, 0.5, "Boundary (a):", adj = 0)
text(0.7, 0.5, a, adj = 0)
text(0.2, 0.4, "Start point (z):", adj = 0)
text(0.7, 0.4, paste0(z_prop, " (", round(z, 2), ")"), adj = 0)
text(0.2, 0.3, "Noise (s):", adj = 0)
text(0.7, 0.3, s, adj = 0)
text(0.2, 0.2, "Non-decision (Ter):", adj = 0)
text(0.7, 0.2, Ter, adj = 0)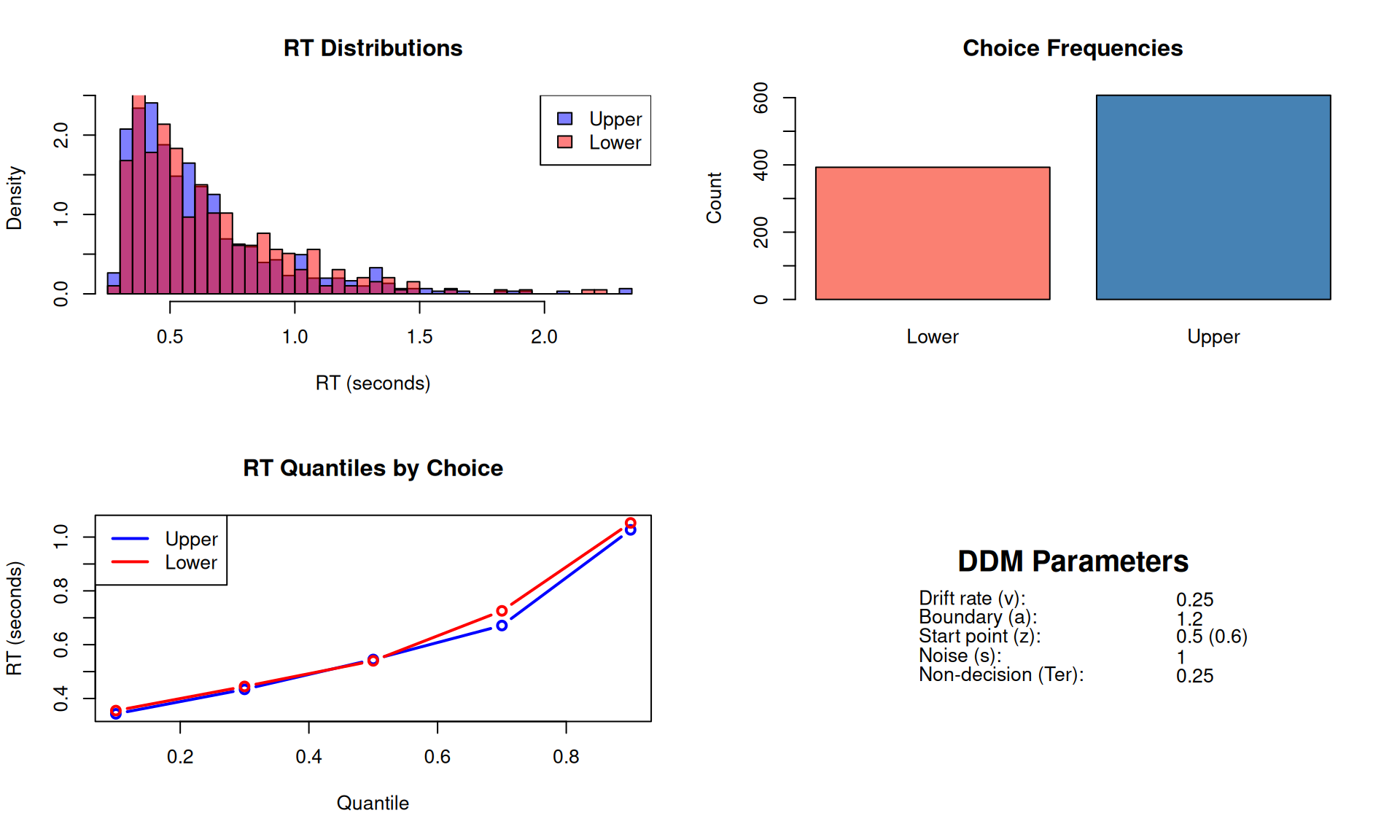
1. Random walk → pure noise, no direction 2. Add drift → systematic tendency 3. Add boundaries → decisions emerge 4. Add non-decision time → realistic RTs 5. Complete DDM → accounts for accuracy AND RT